
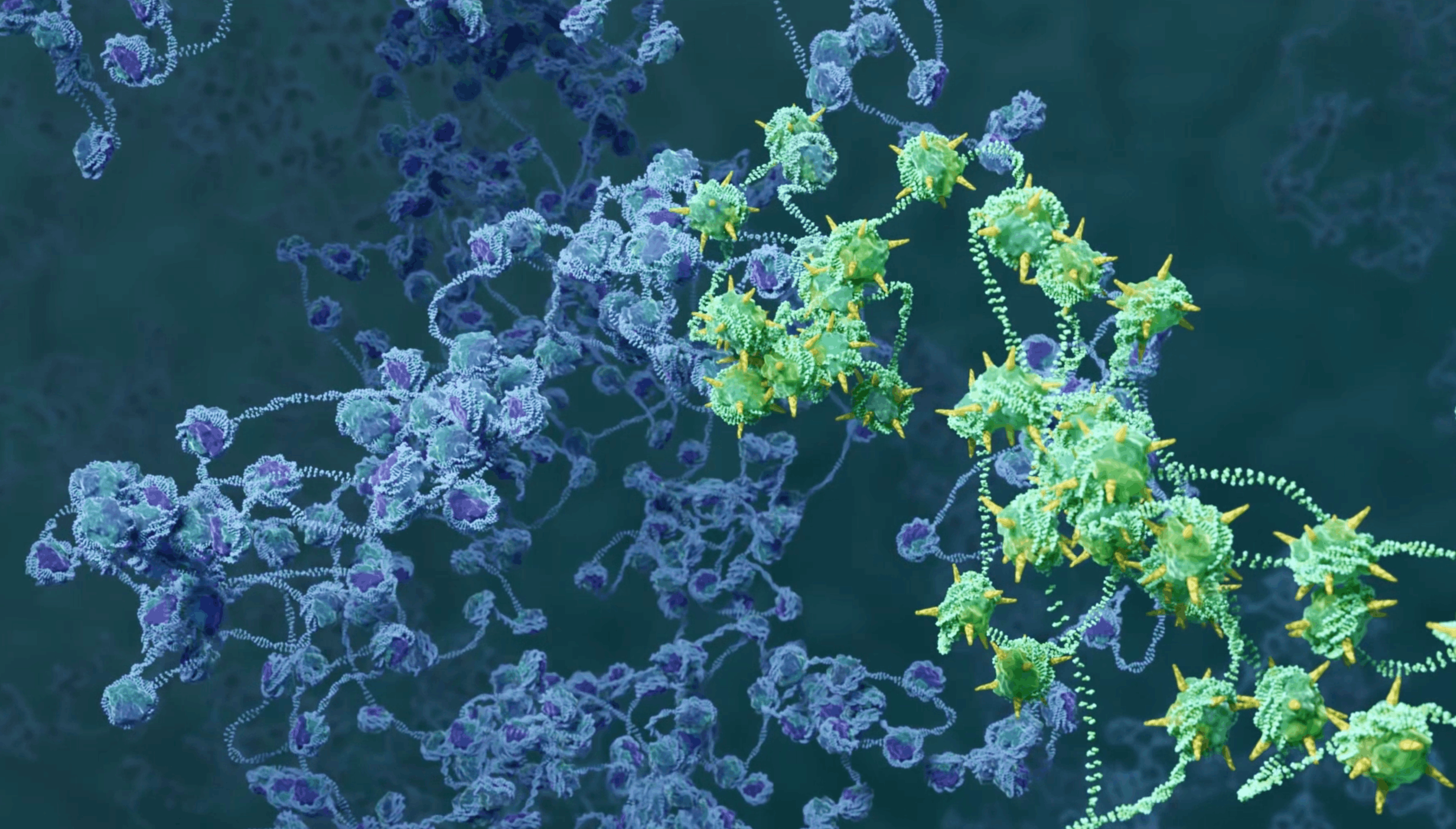
Transient histone deacetylase inhibition induces cellular memory of gene expression and 3D genome folding. Nat Genetics, in press.
Paldi, F., Szalay, M.F., Dufau, S., Di Stefano, M., Reboul, H., Jose, D., Bantignies, F., and Cavalli, G. (2026). 10.1038/s41588-025-02489-4
5 February 2026

Cell competition overcomes host tissue resistance to unleash tumour growth2 in a Drosophila brain cancer model
Cédric Maurange and Pauline Spéder
9 December 2025

Uniform dynamics of cohesin-mediated loop extrusion in living human cells | Nature Genetics
Christophe Zimmer, Edouard Bertrand
27 November 2025
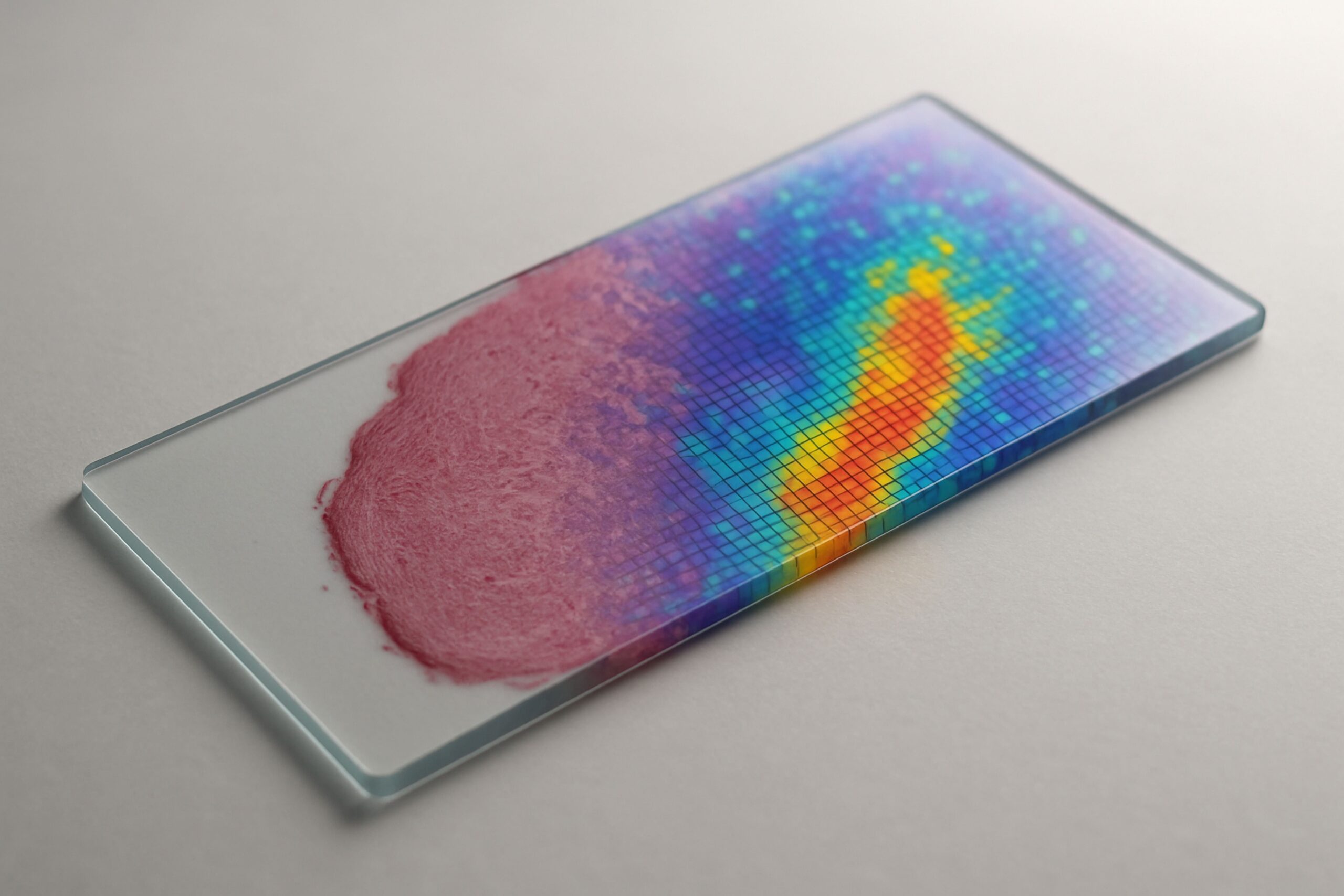
RNA2seg: a generalist model for cell segmentation in image-based spatial transcriptomics
Thomas Walter, Florian Mueller
14 November 2025
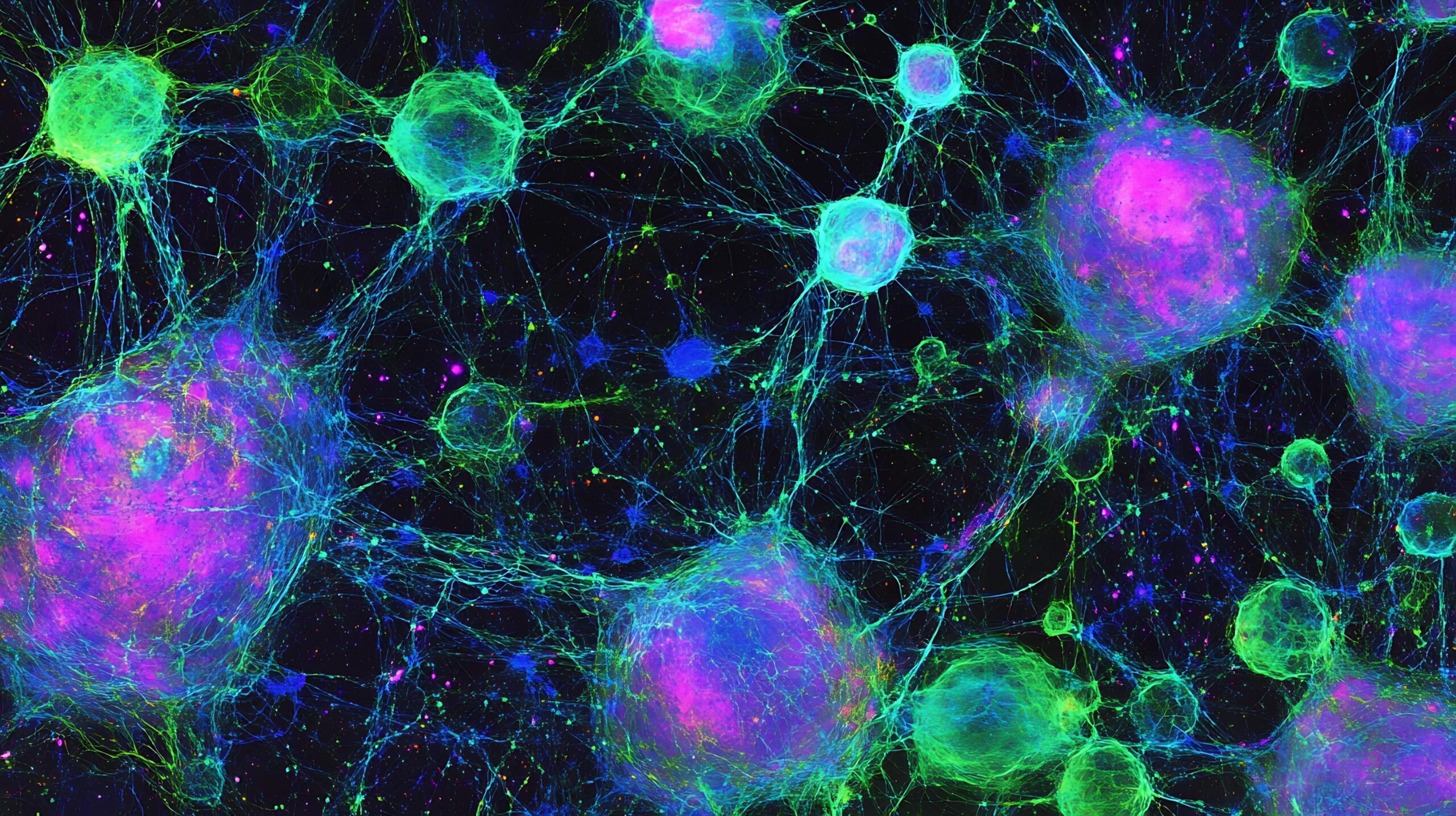
MIPHEI-ViT: Multiplex Immunofluorescence Prediction from H&E Images using ViT Foundation Models
Thomas Walter
14 November 2025

A Deep Learning approach for time-consistent cell cyclephase prediction from microscopy data
Thomas Walter
14 November 2025

STORIES: learning cell fate landscapes from spatial transcriptomics using optimal transport
Laura Cantini
14 November 2025
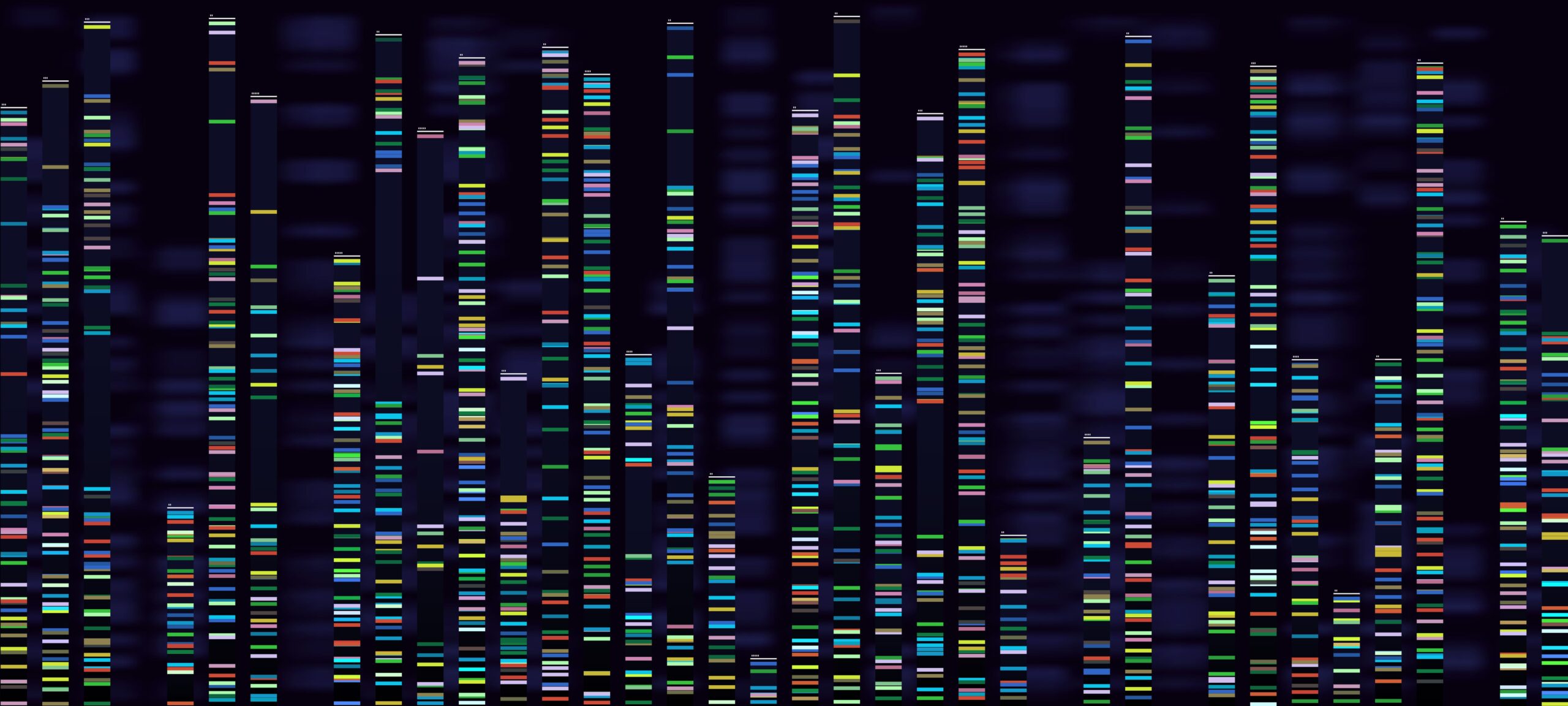
HIRA defines early replication initiation zones independently of their genome compartment
Geneviève Almouzni
14 November 2025

Loop extrusion provides mechanical robustness to chromatin
Hossein Salari, Daniel Jost
14 November 2025

FAIR sharing of Chromatin Tracing datasets using the newly developed 4DN FISH Omics Format
M. Nollmann
14 November 2025
Pas d’actualités